Antimicrobial resistance is one of the greatest challenges to global health. Researchers from the Department of “Epidemiology and Ecology of Antimicrobial Resistance” at the Helmholtz Institute for One Health (HIOH), led by Prof. Dr. Katharina Schaufler, PhD, have now uncovered a previously hidden link between antibiotic resistance and virulence in Klebsiella pneumoniae using the functional genomics method TraDIS (Transposon-directed insertion-site sequencing). Virulence describes the extent to which a pathogen can cause disease. The findings were published in the international journal Clinical Microbiology and Infection.
AMR: a global health challenge
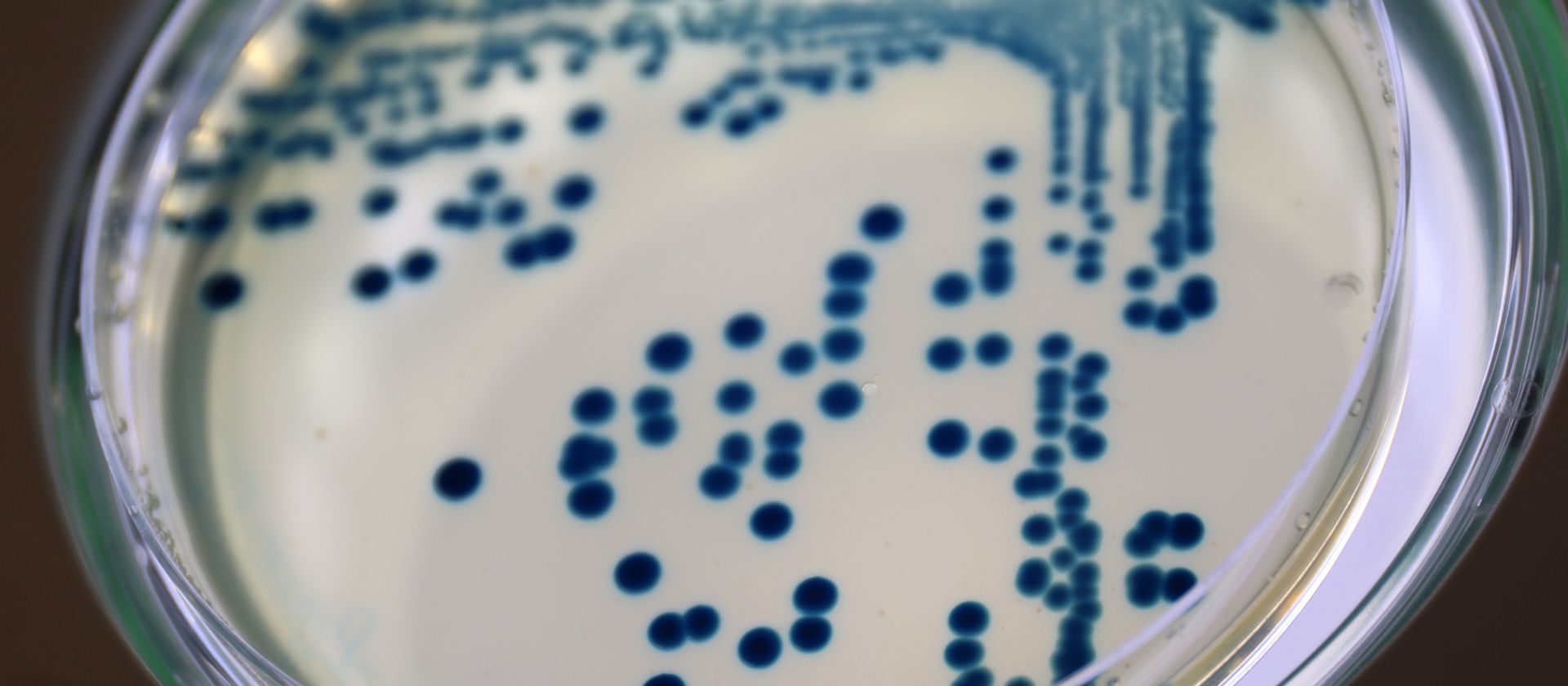
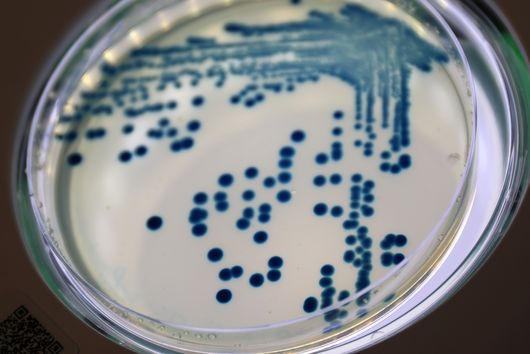
Klebsiella pneumoniae is one of the most important causes of severe infections worldwide, particularly in healthcare settings (so-called nosocomial infections). From a One Health perspective, the bacterium is of particular relevance, as it occurs in the environment and can cause infections in humans and animals alike. Especially problematic are strains that are resistant to multiple antibiotics. In such cases, only very limited treatment options remain.
The reserve antibiotic cefiderocol is considered a promising treatment option against these resistant bacteria. It acts like a “Trojan horse”: it disguises itself as iron, which is essential for bacterial survival, is actively transported into the cell, and then kills the bacteria. However, in recent years, increasing reports have shown that resistance is now emerging even against cefiderocol.

































![[Translate to English:] [Translate to English:]](/fileadmin/HIOH/__processed__/8/3/csm_Lab_Eroeffnung_Buake1_c937de8203.jpeg)